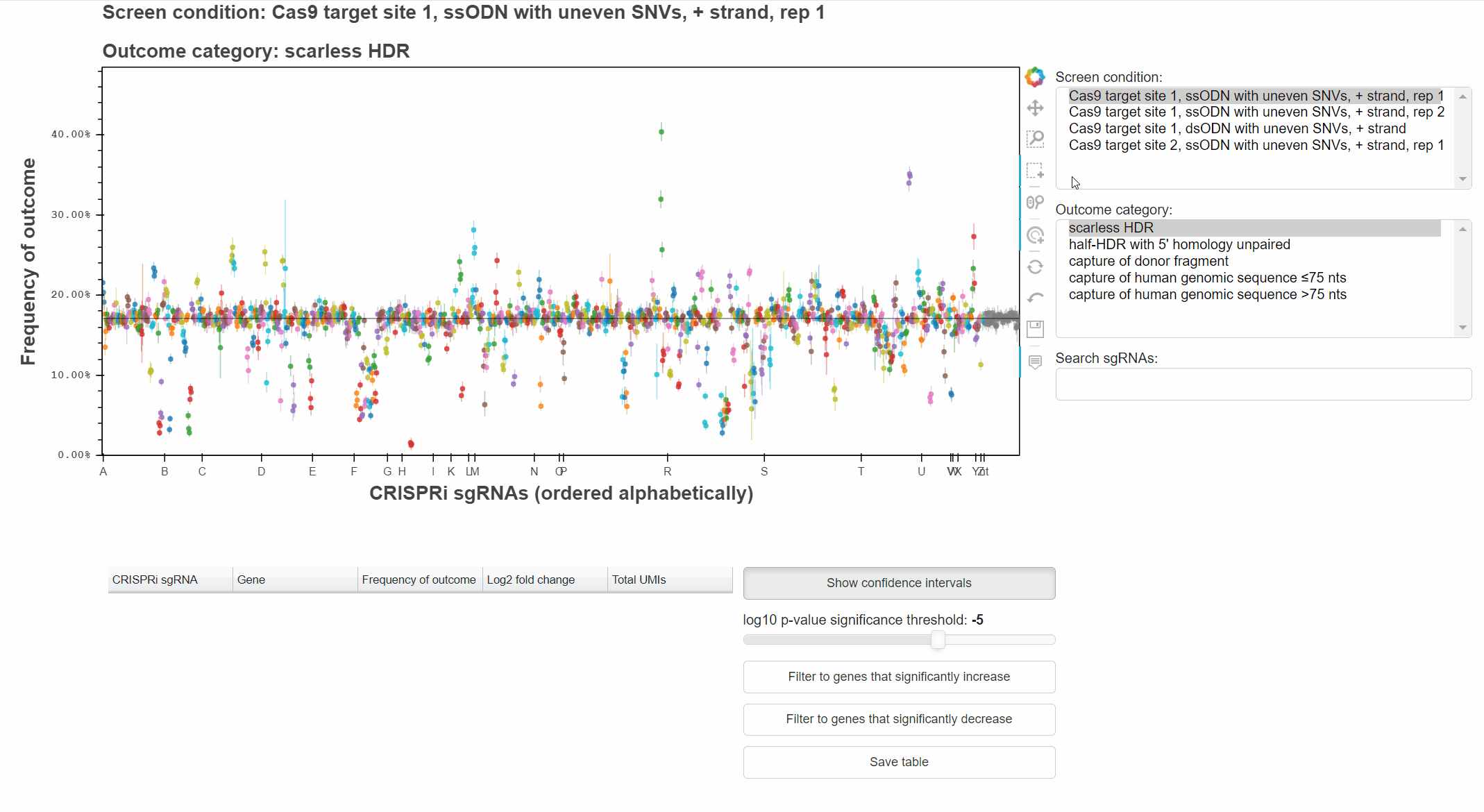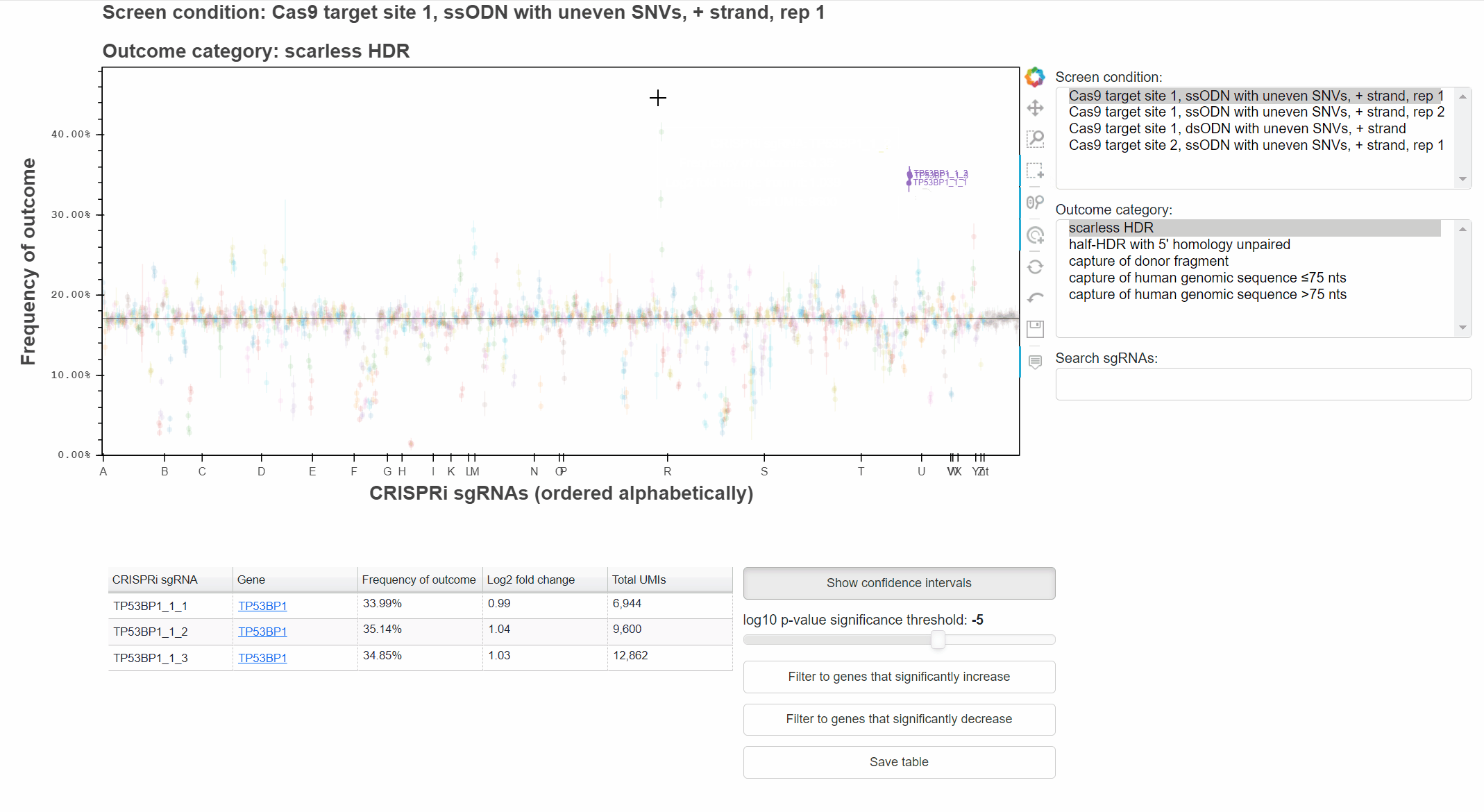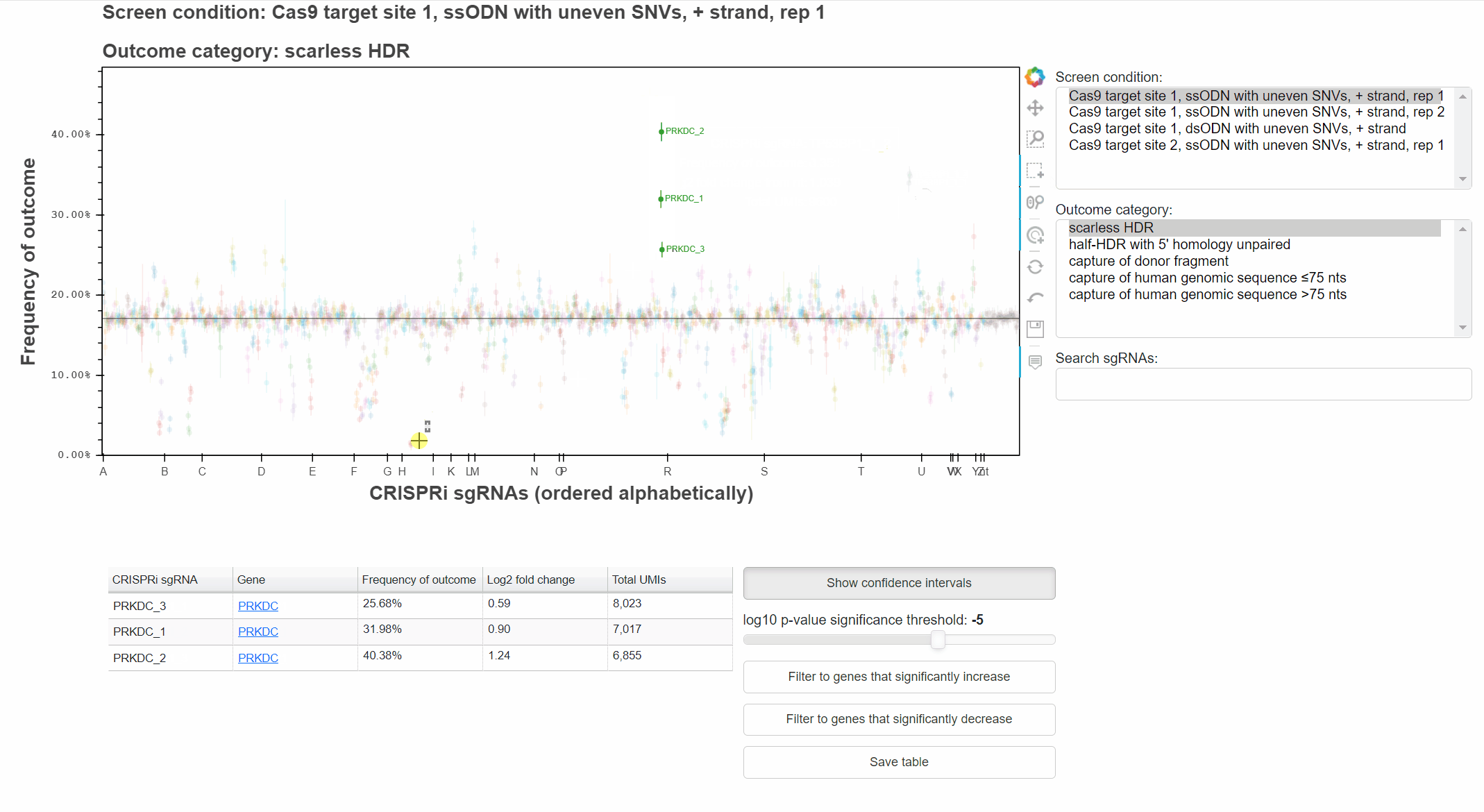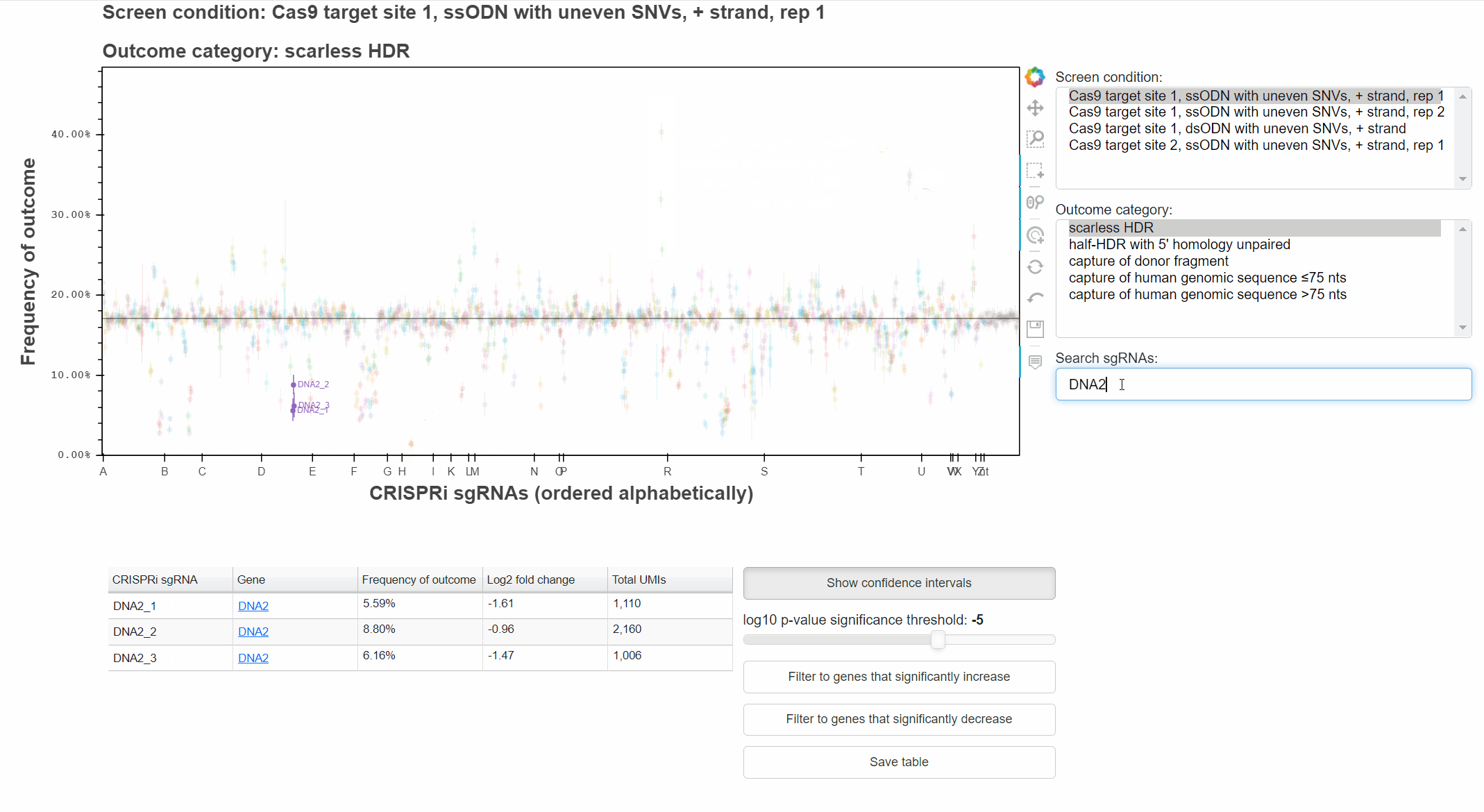
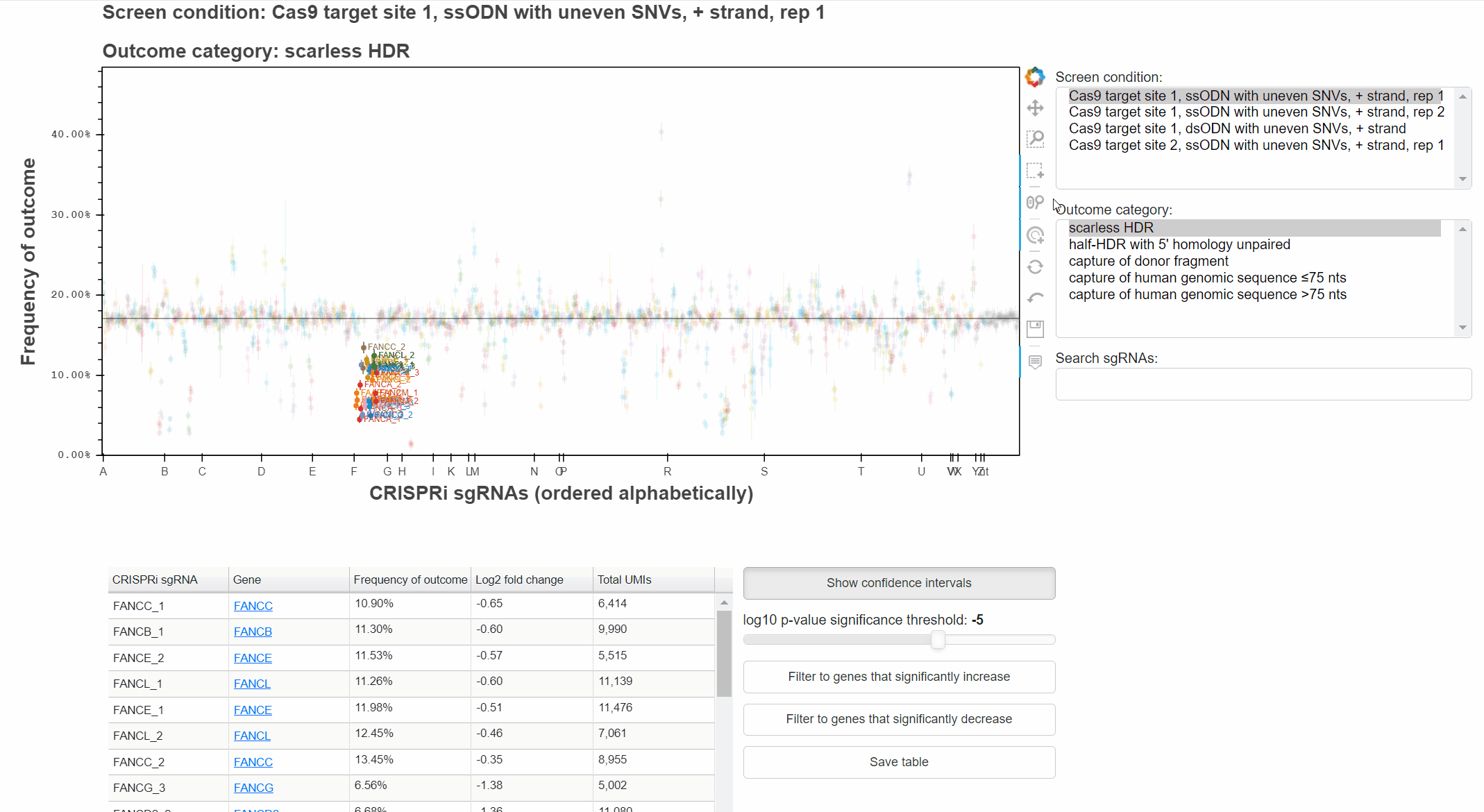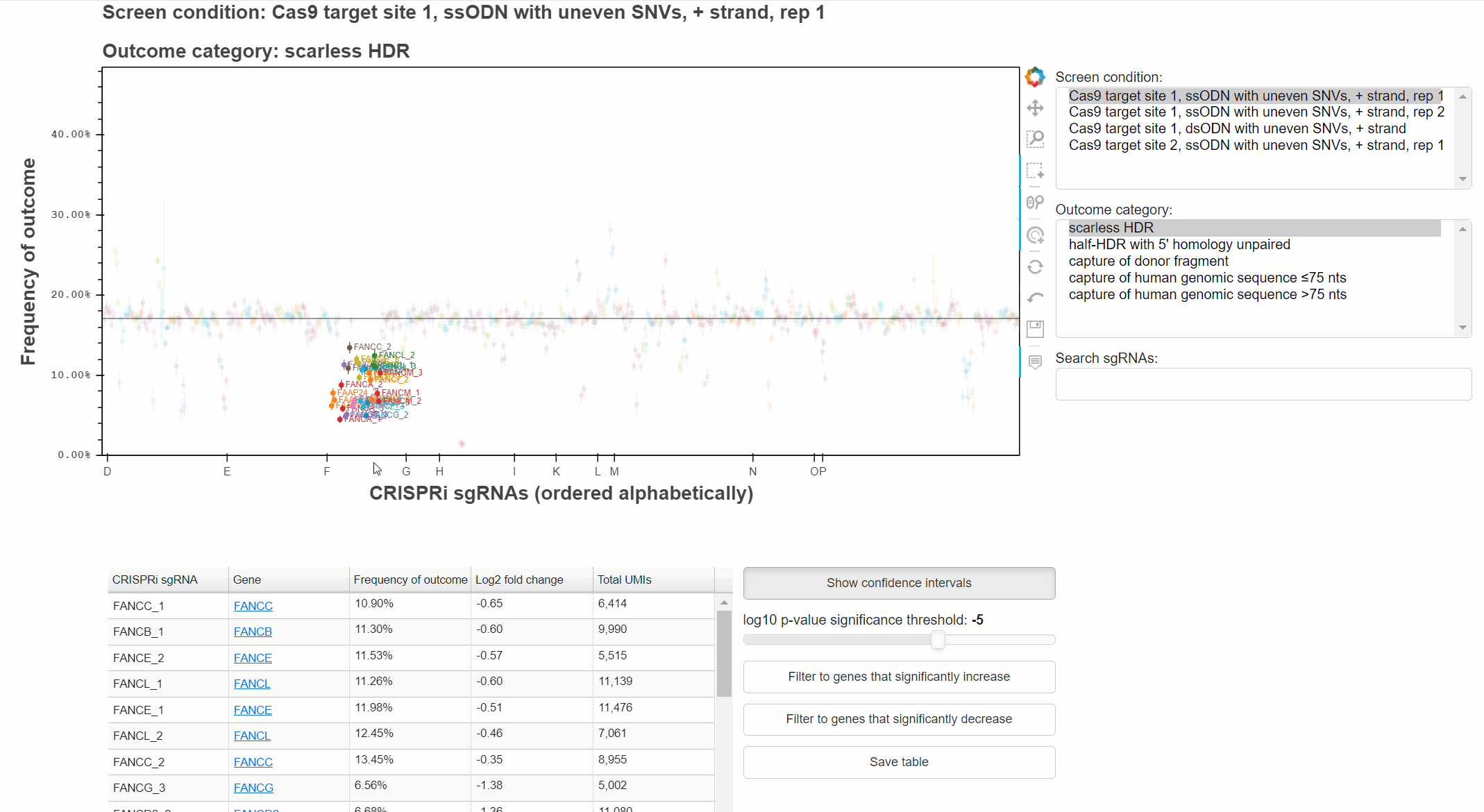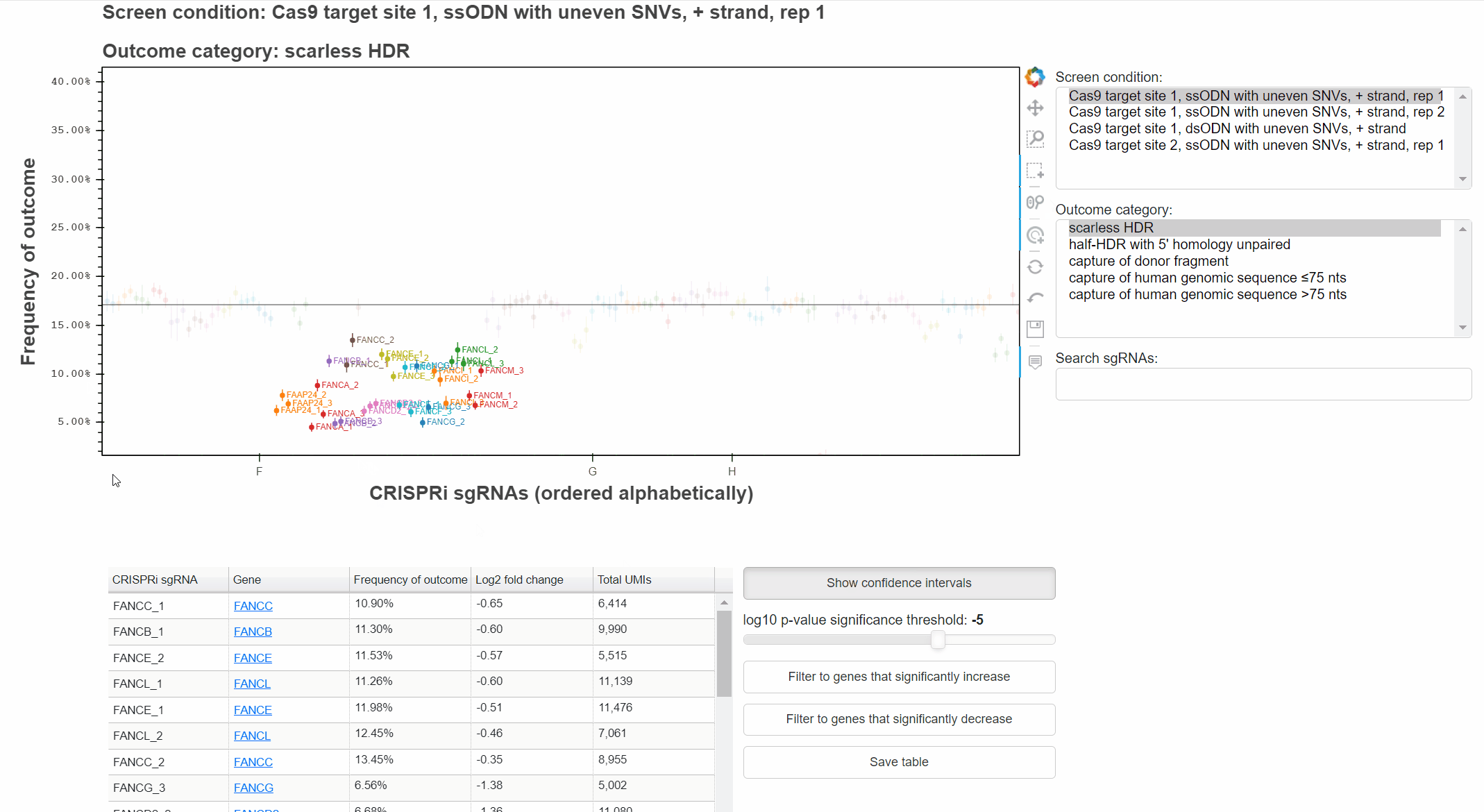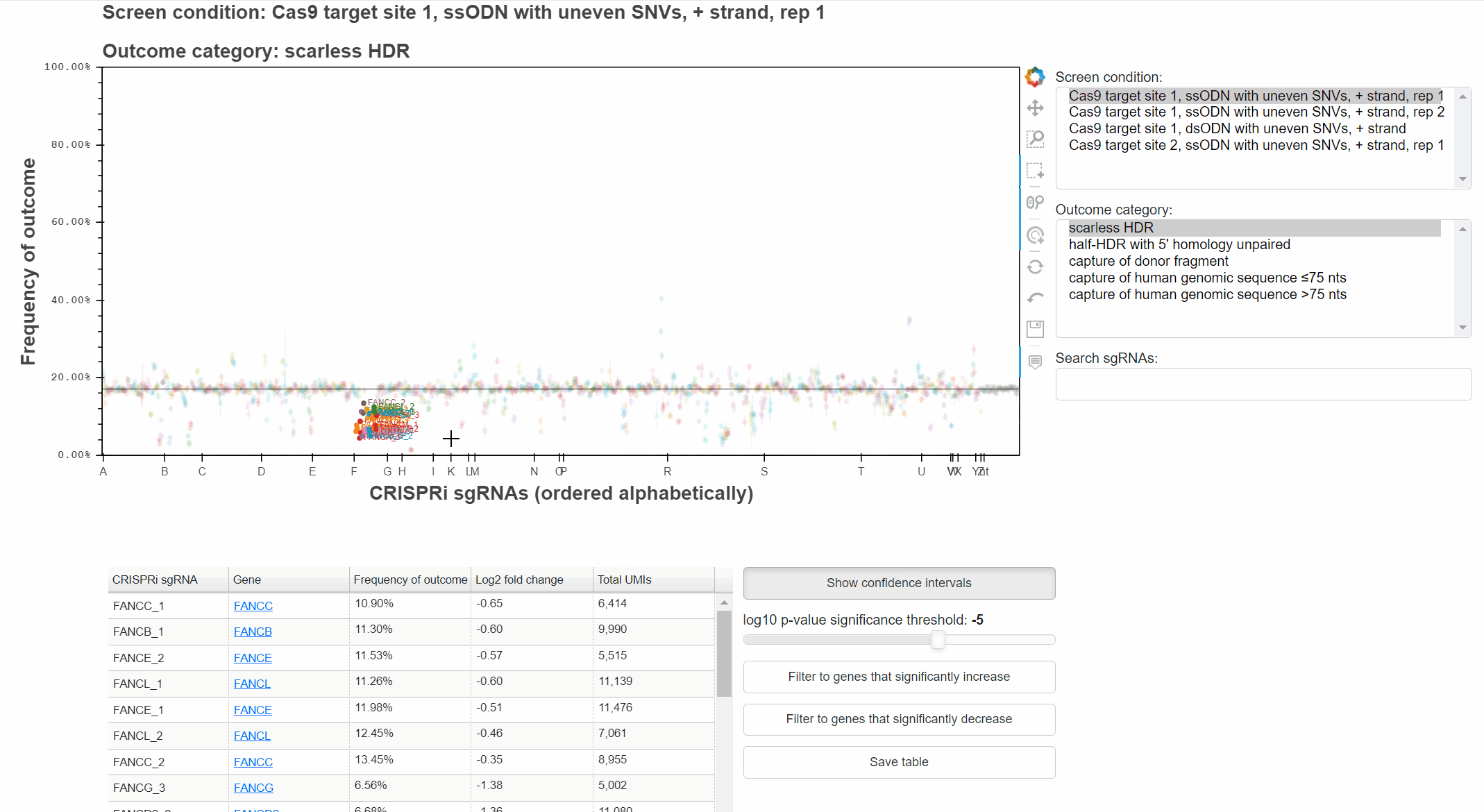
sgRNAs are ordered alphabetically along the x-axis to highlight when multiple sgRNAs targetting the same gene produce consistent phenotypes.
Buttons to the right of the plot toggle different tools for interacting with the plot.
Active tools are marked by a blue bar to the left of the button.
For example, with the "Hover" tool active, hover the cursor over a point to learn which sgRNA it represents.
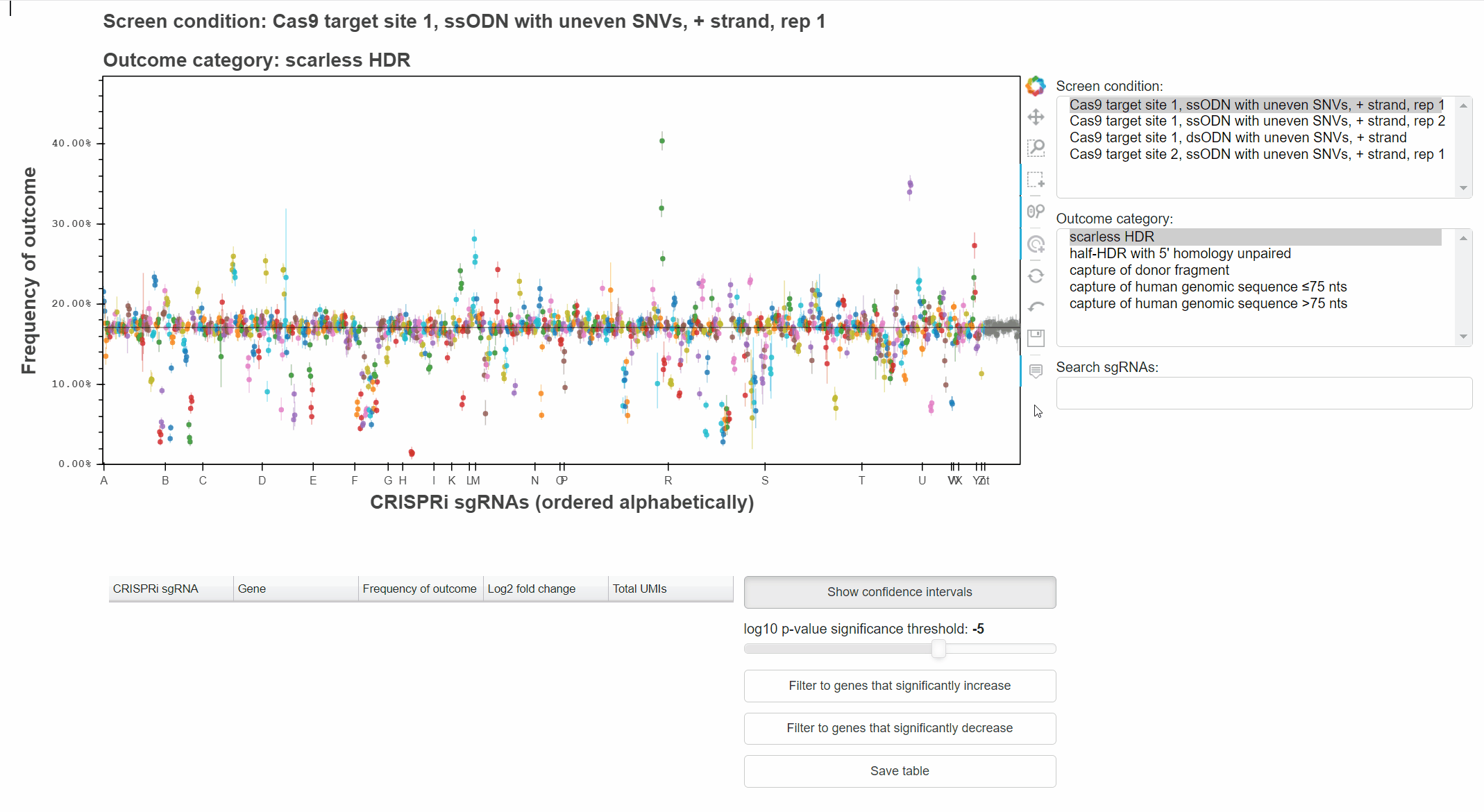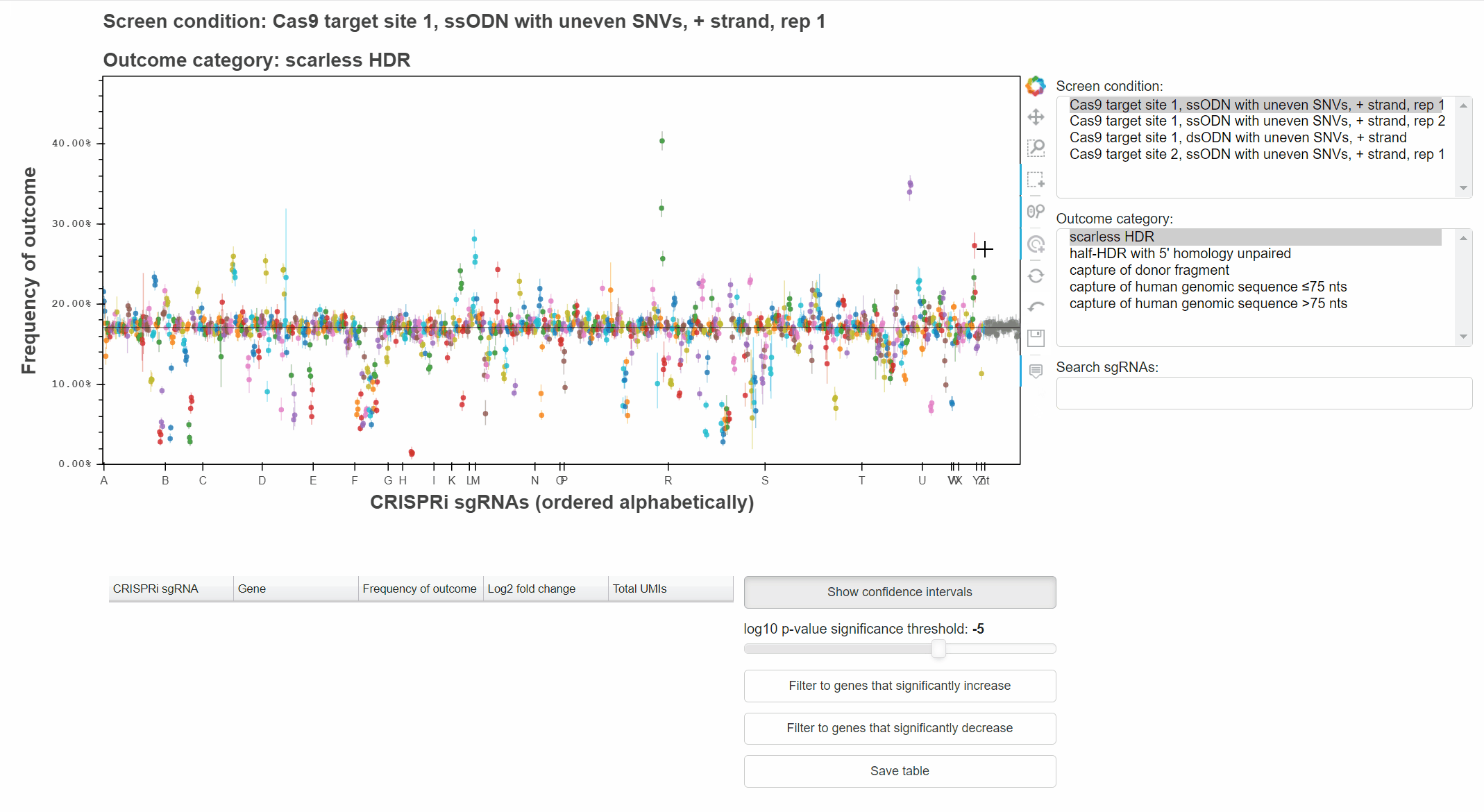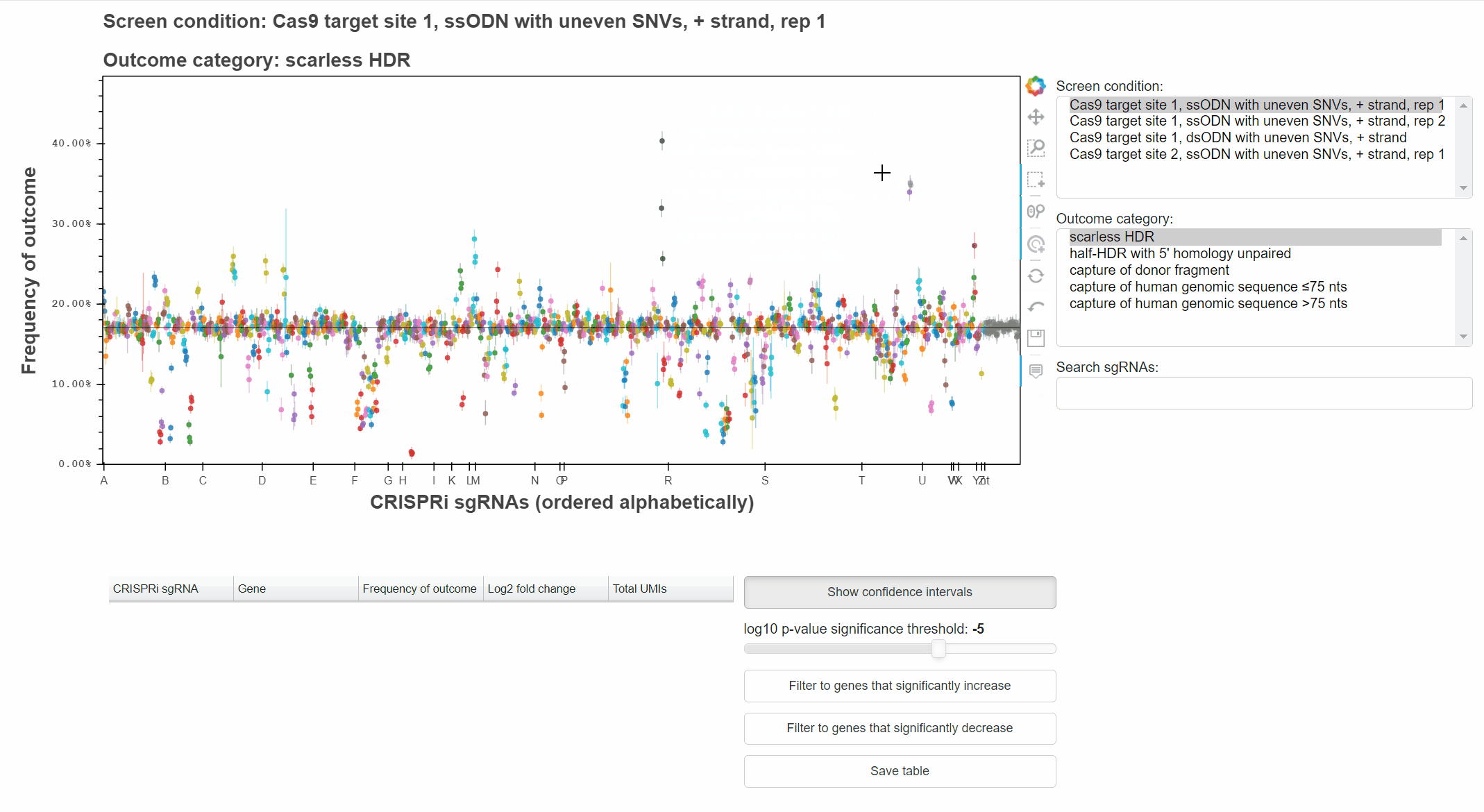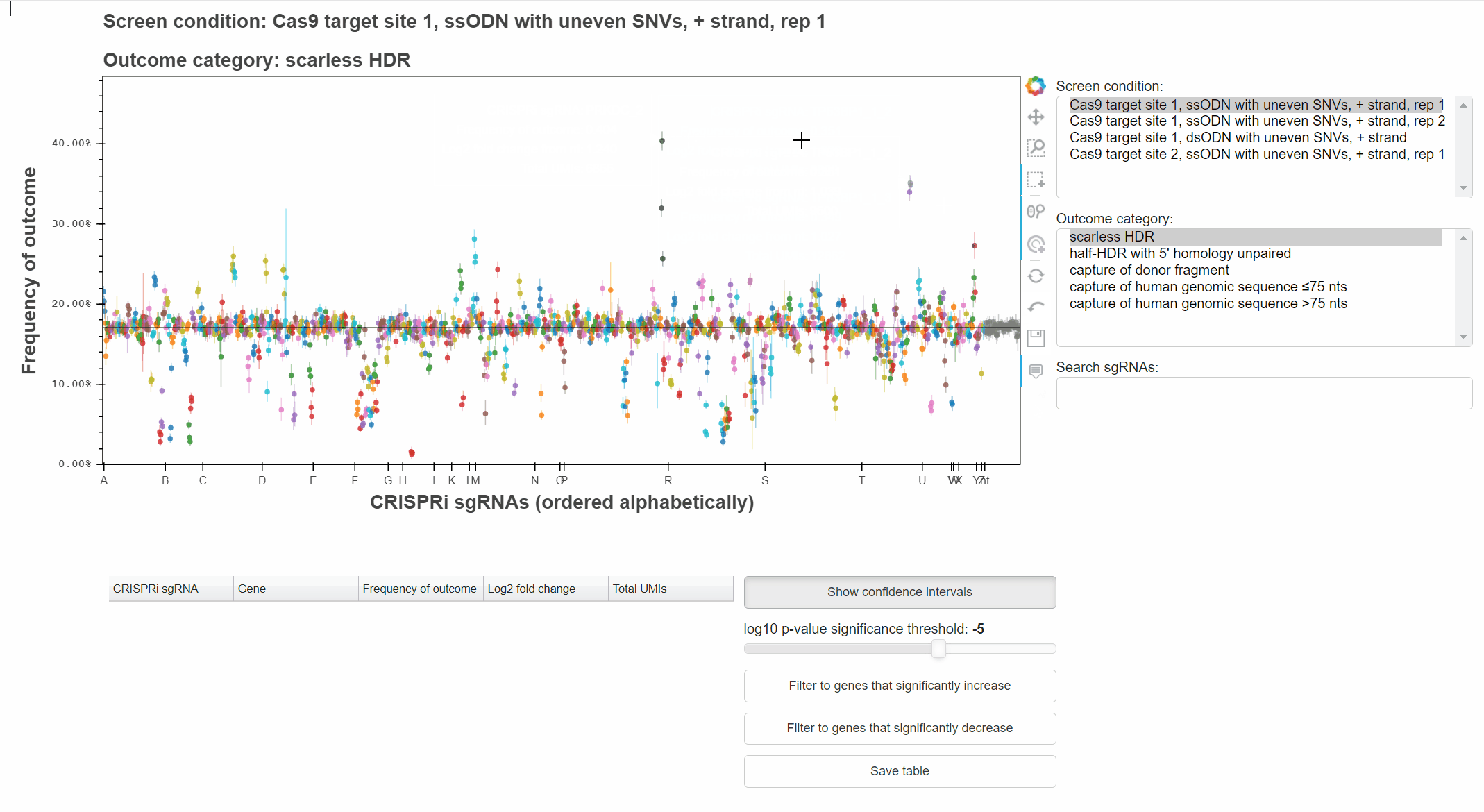
With the "Box Select" tool active, click and drag to select points in a region.
Alternatively, type a query into the search field and hit enter to select matching sgRNAs.
To change the screen or outcome category plotted, click in the relevant menus, or use the up and down arrow keys once a menu is focused.
Highlighting of sgRNAs will persist as the data source is changed, allowing comparison of phenotypes across conditions or outcomes.
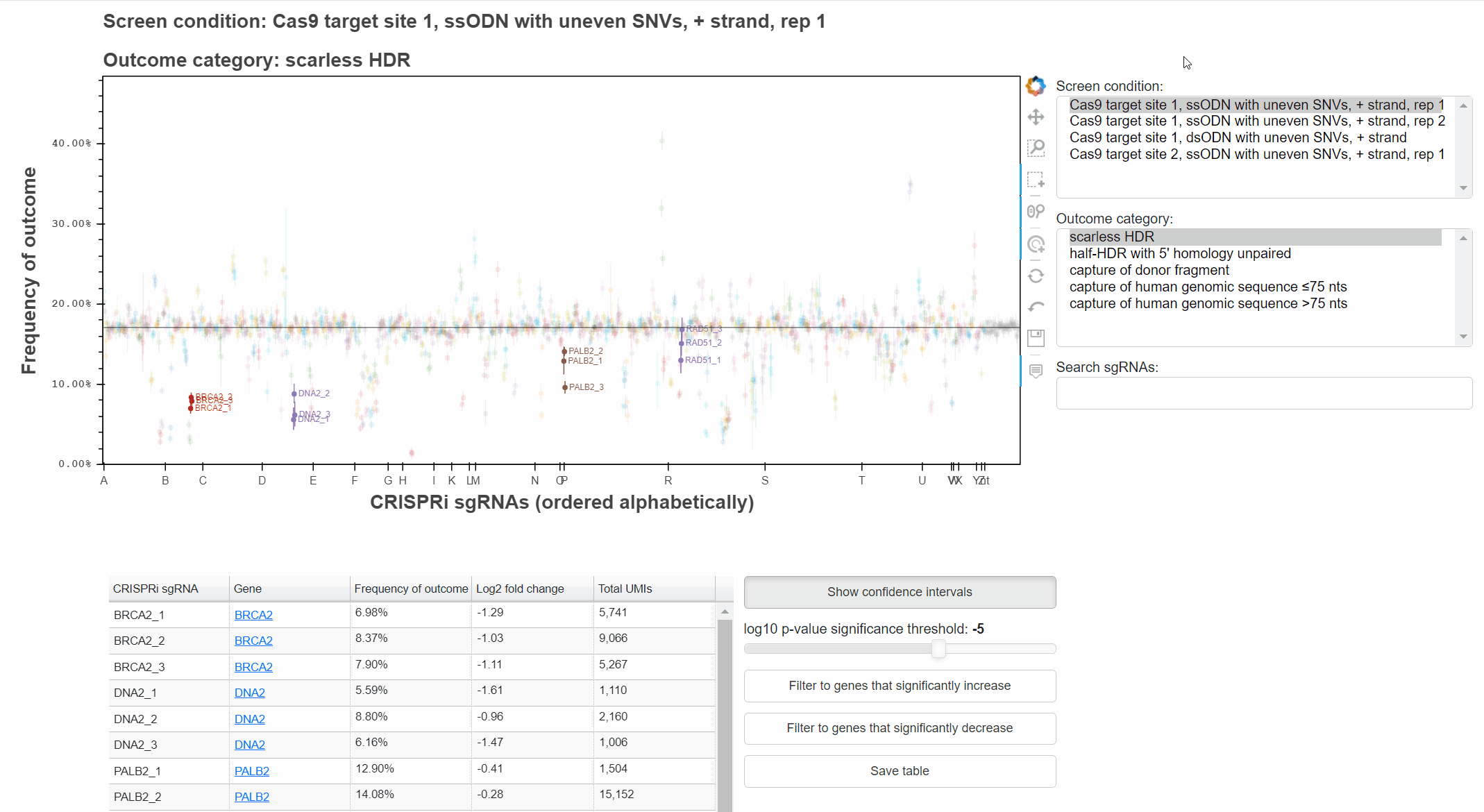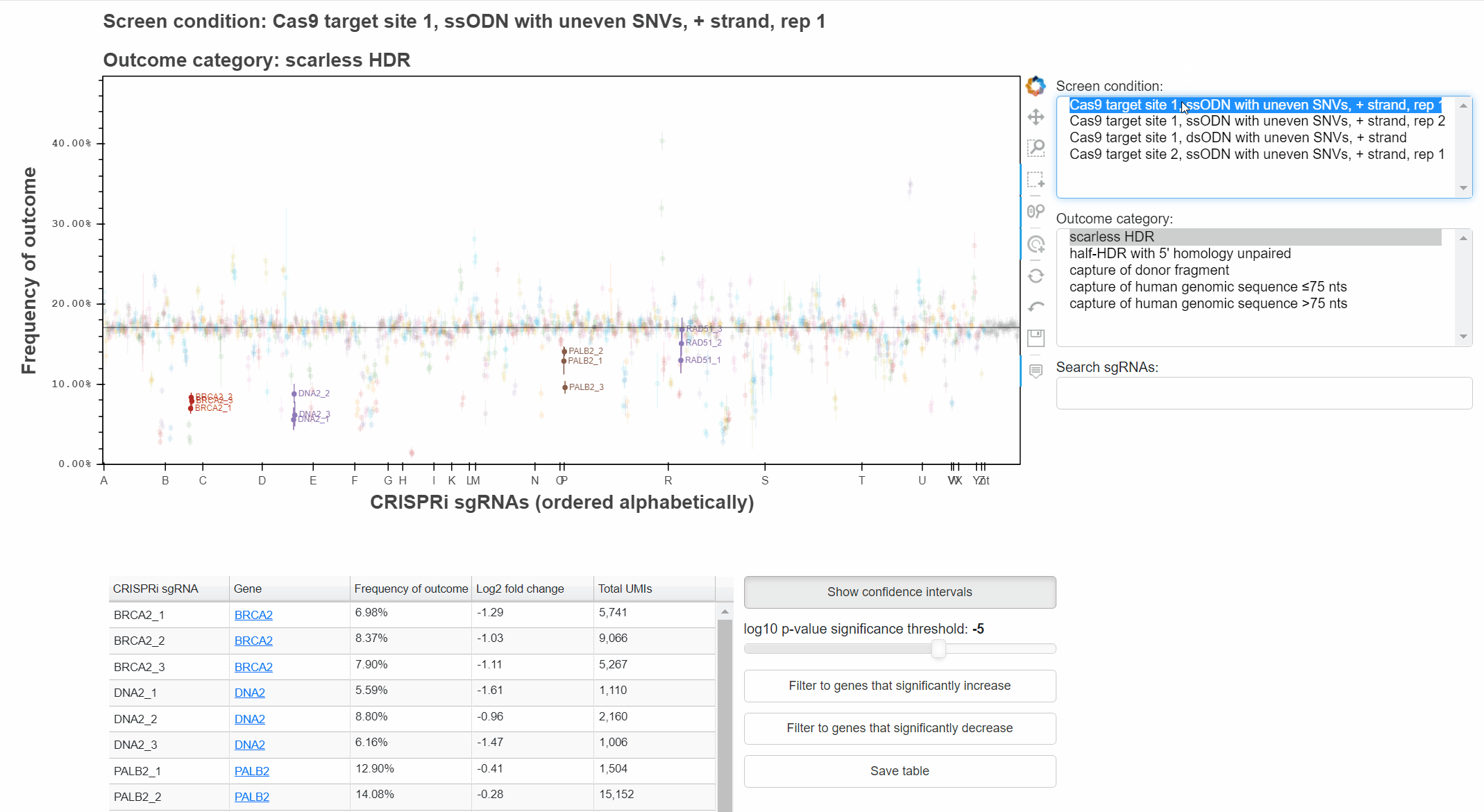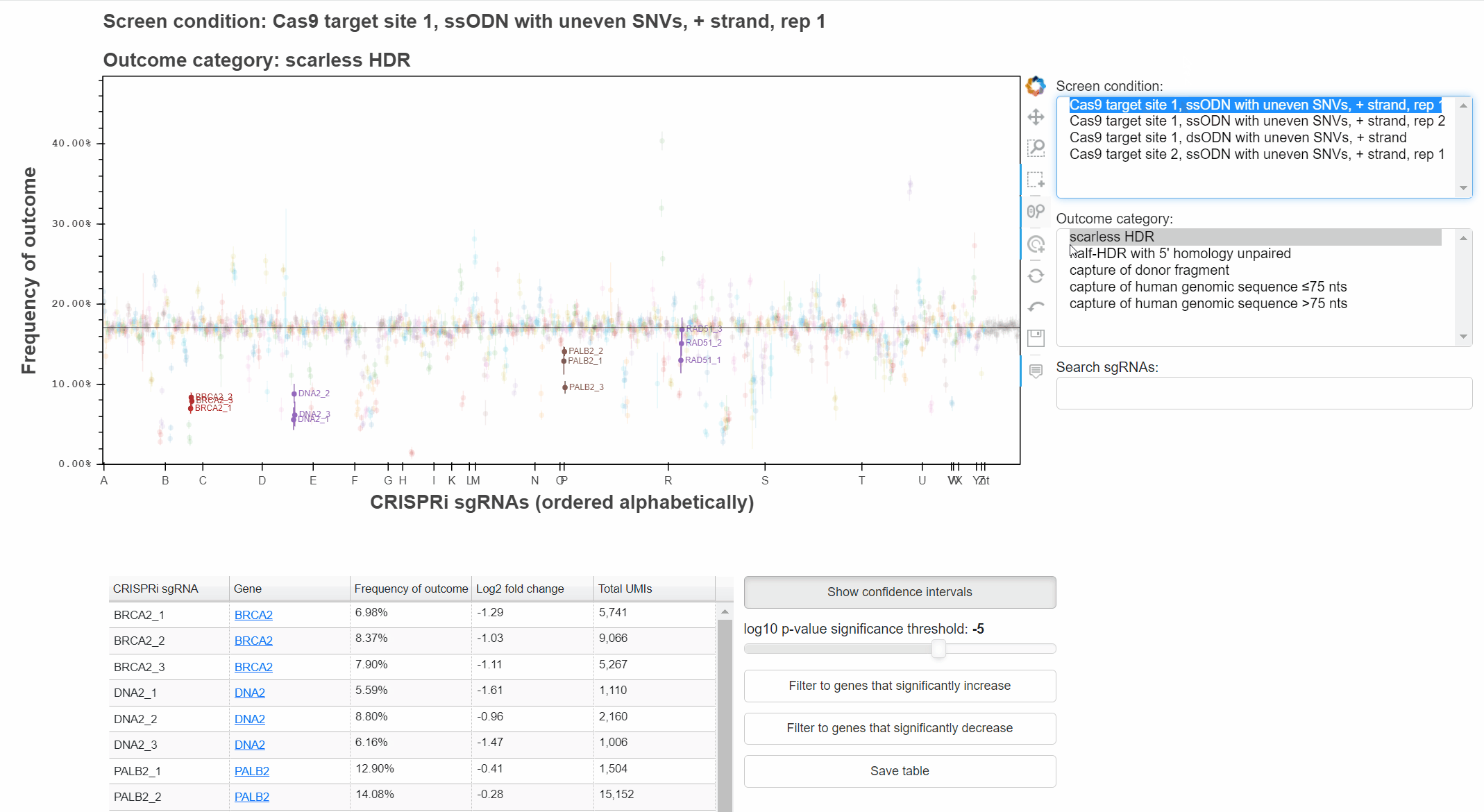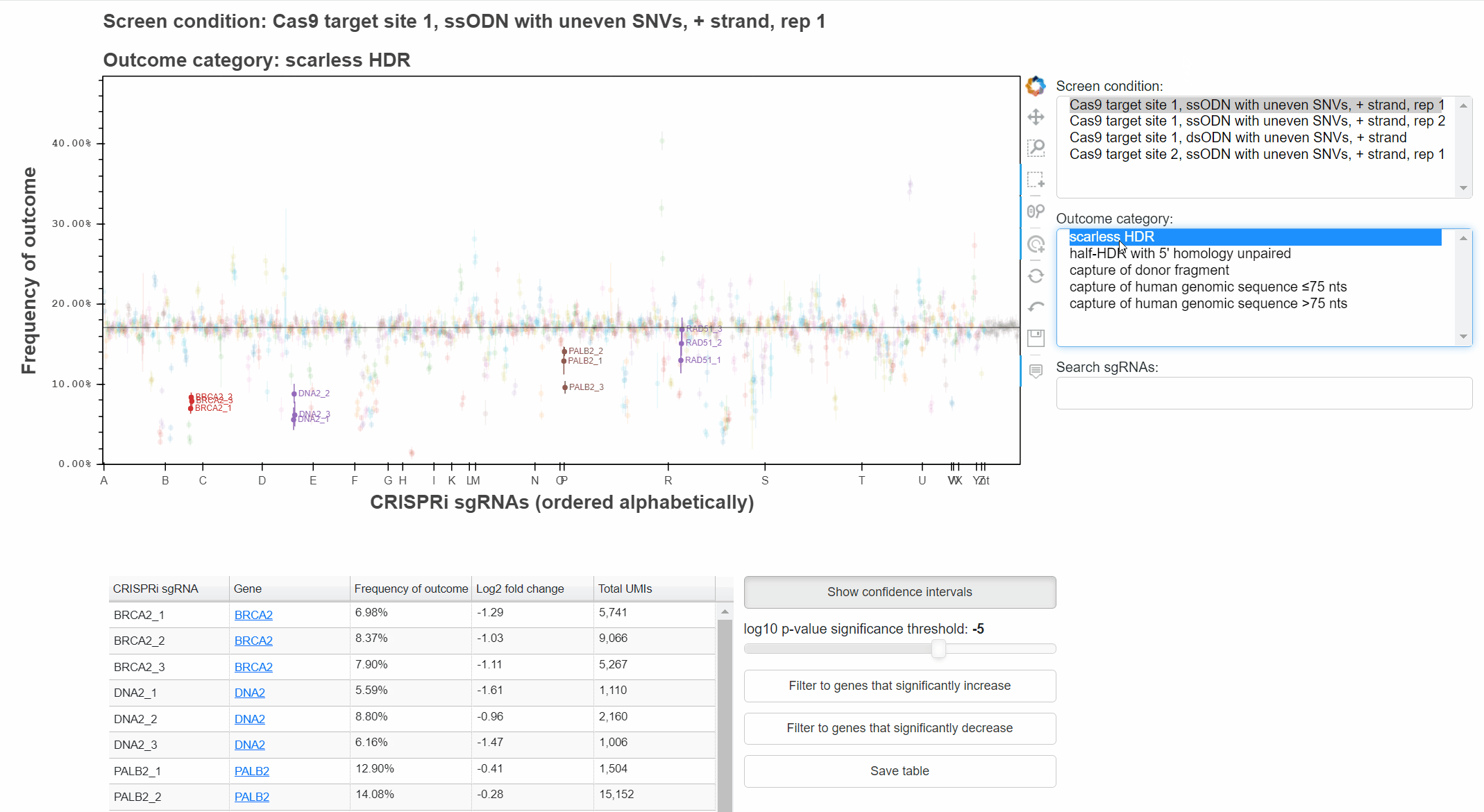
The "Show confidence intervals" button toggles 95% binomial confidence intervals around each sgRNA's outcome frequency.
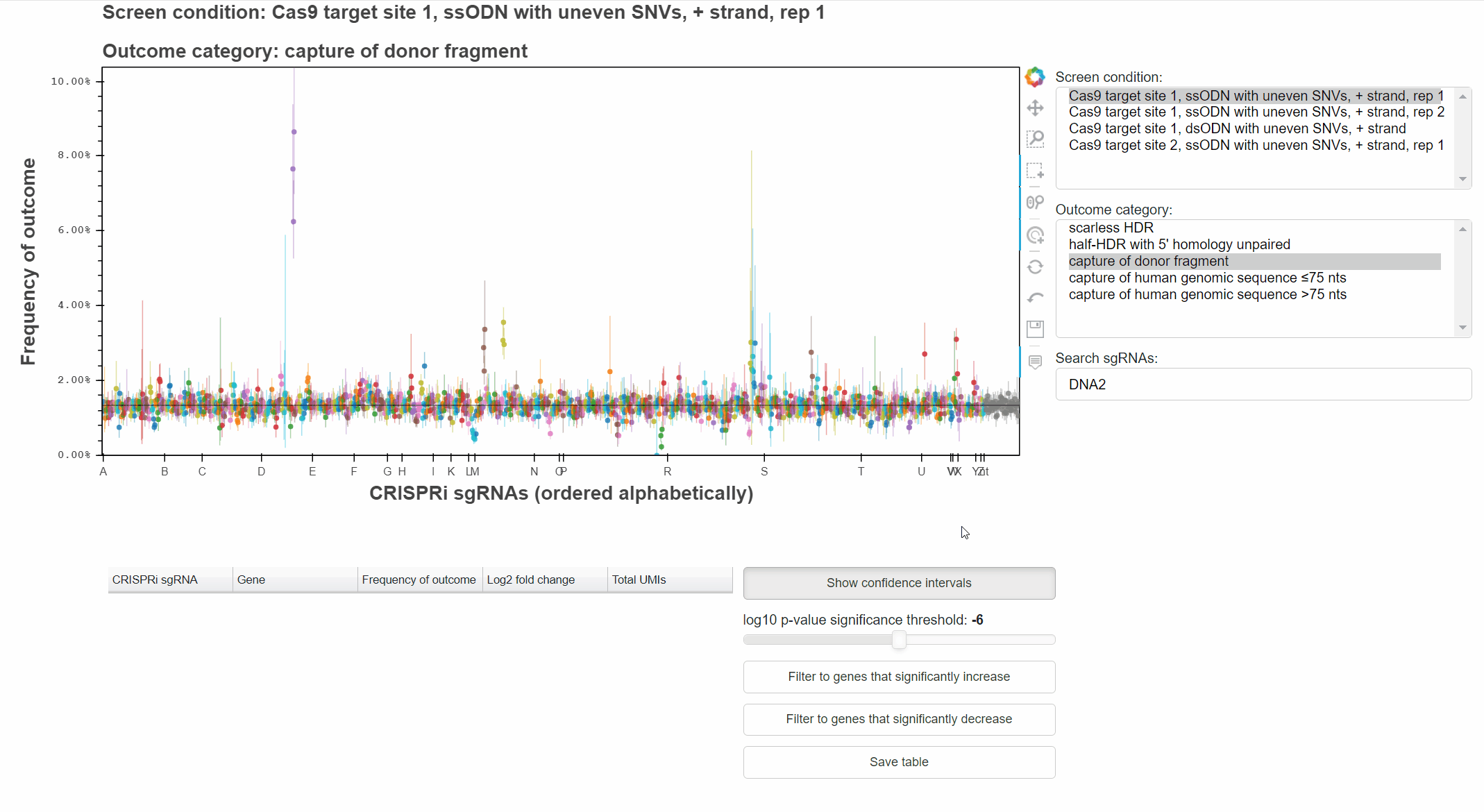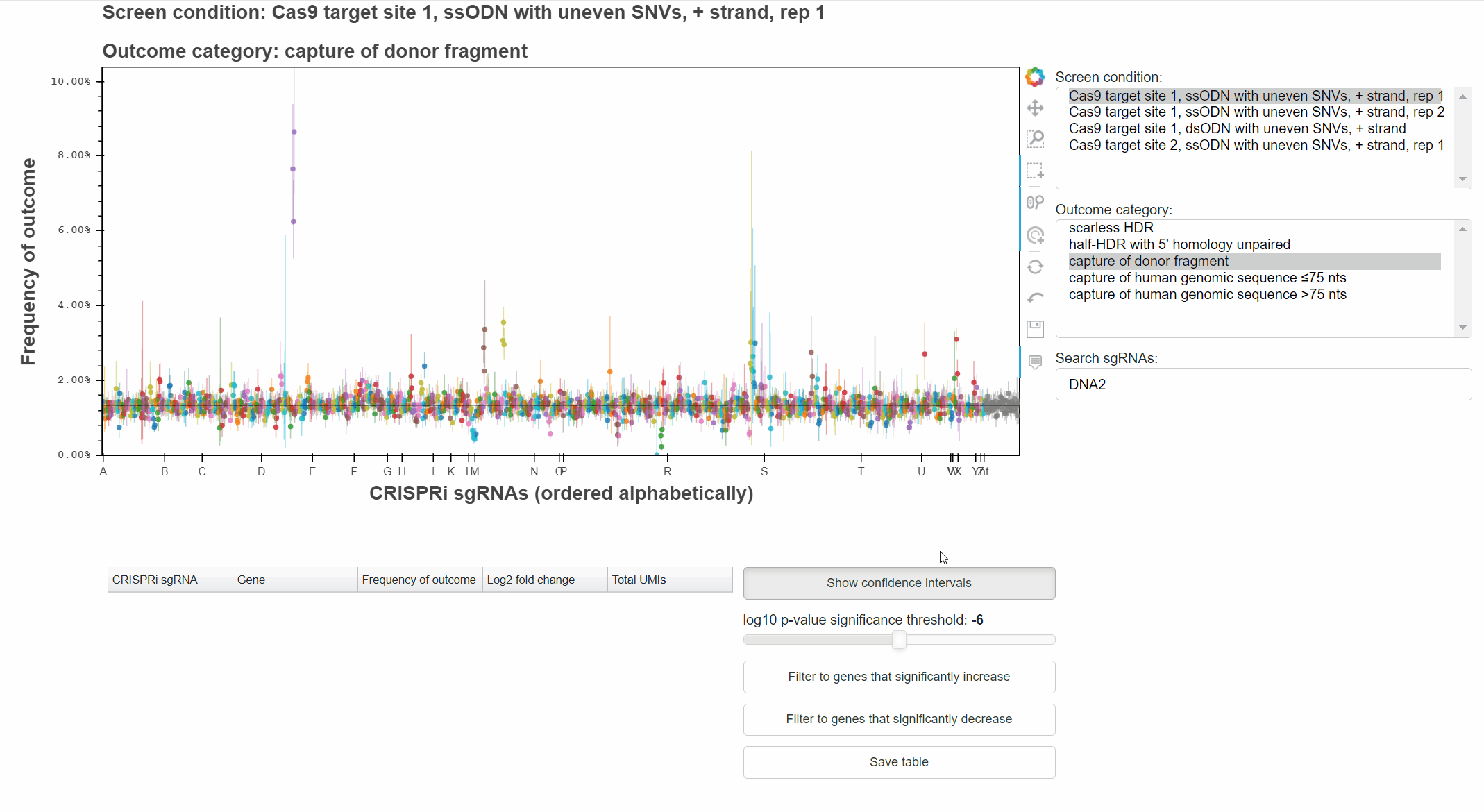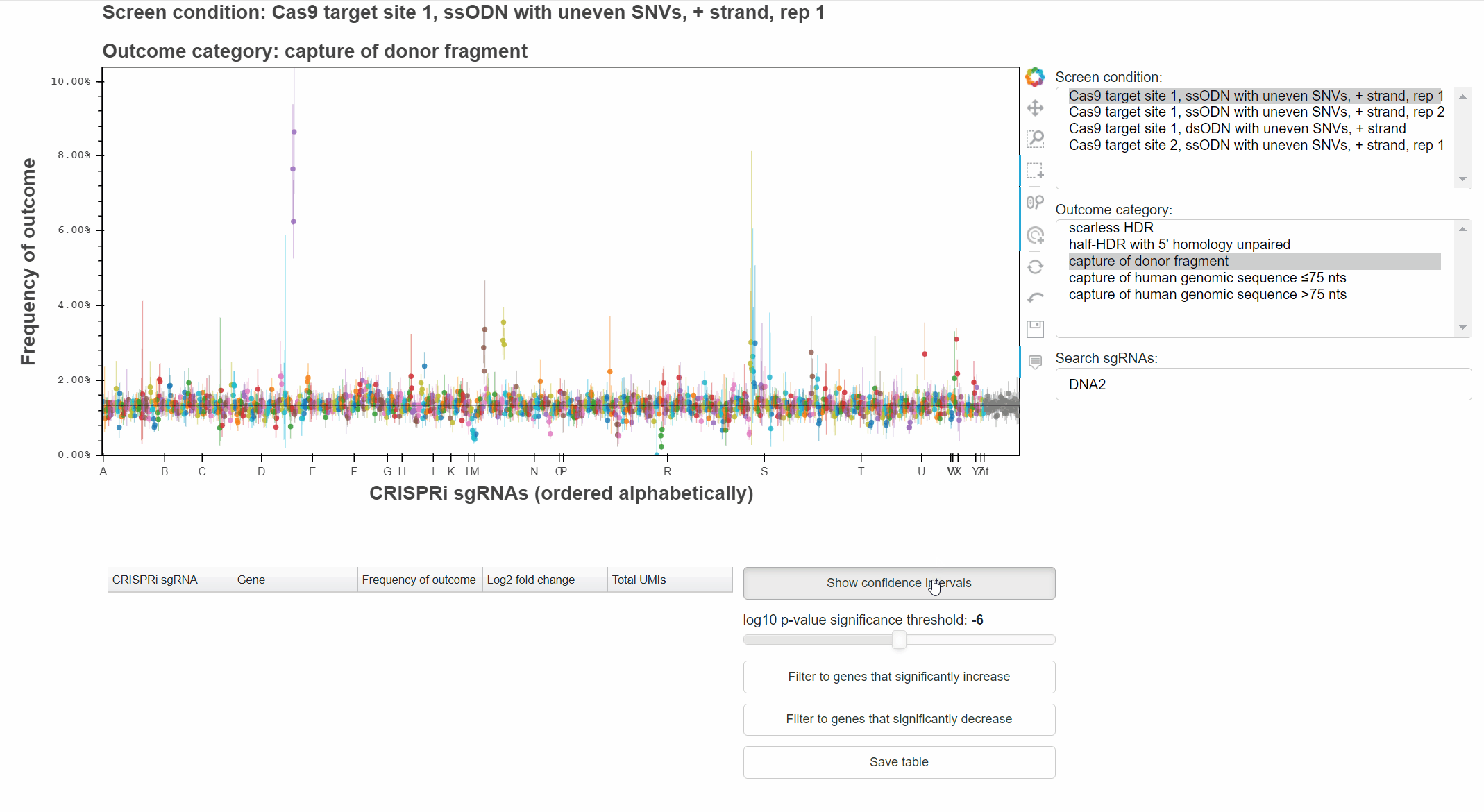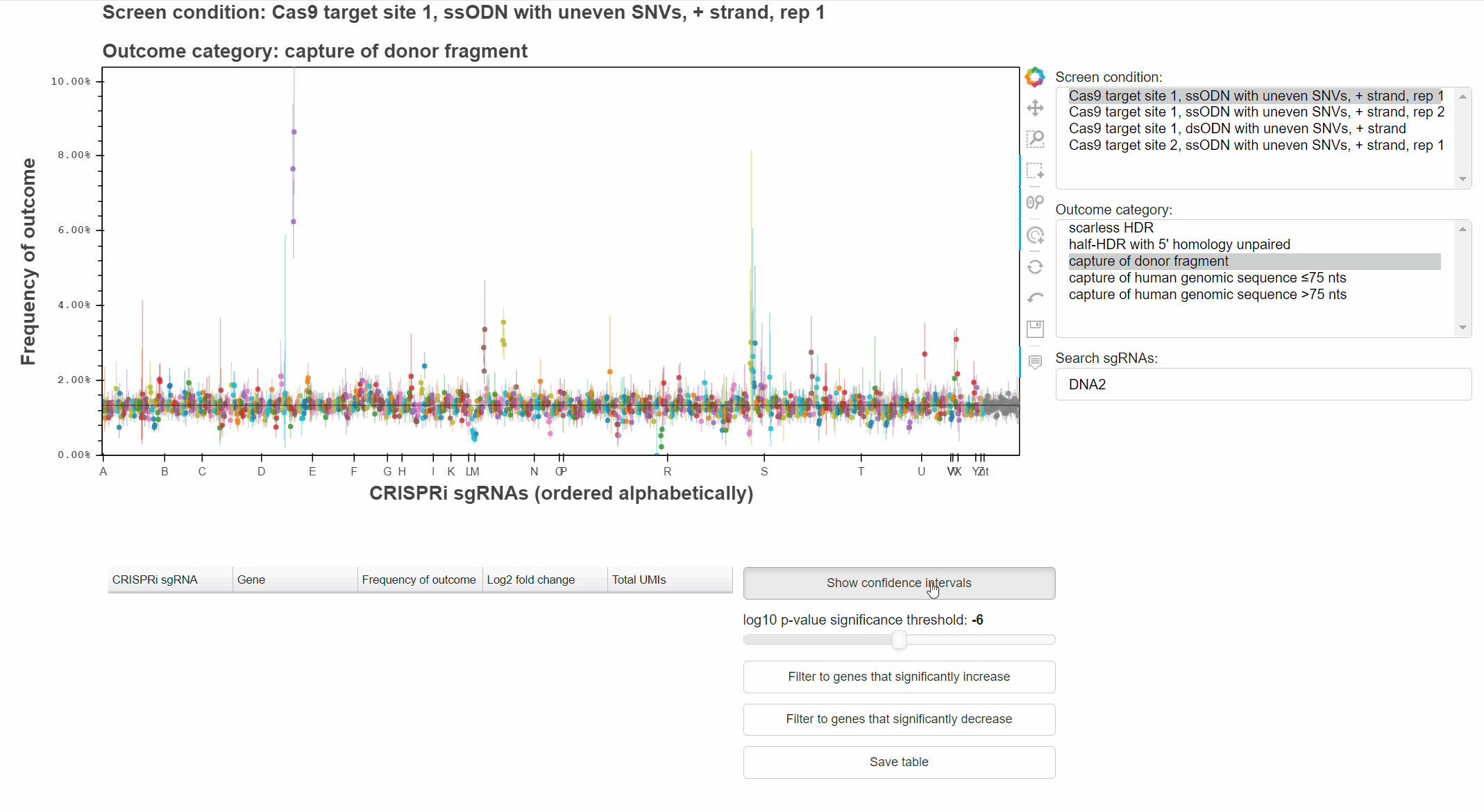
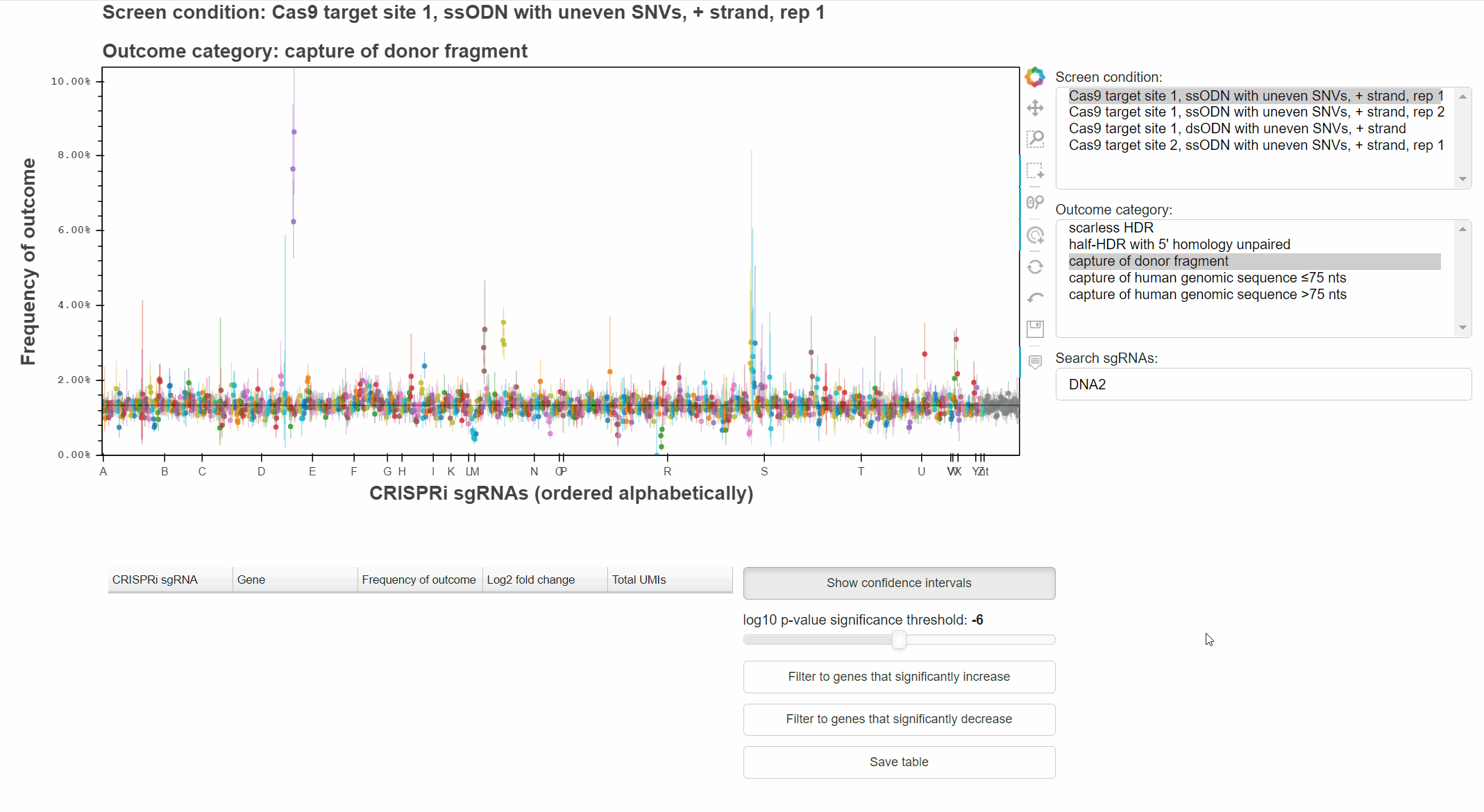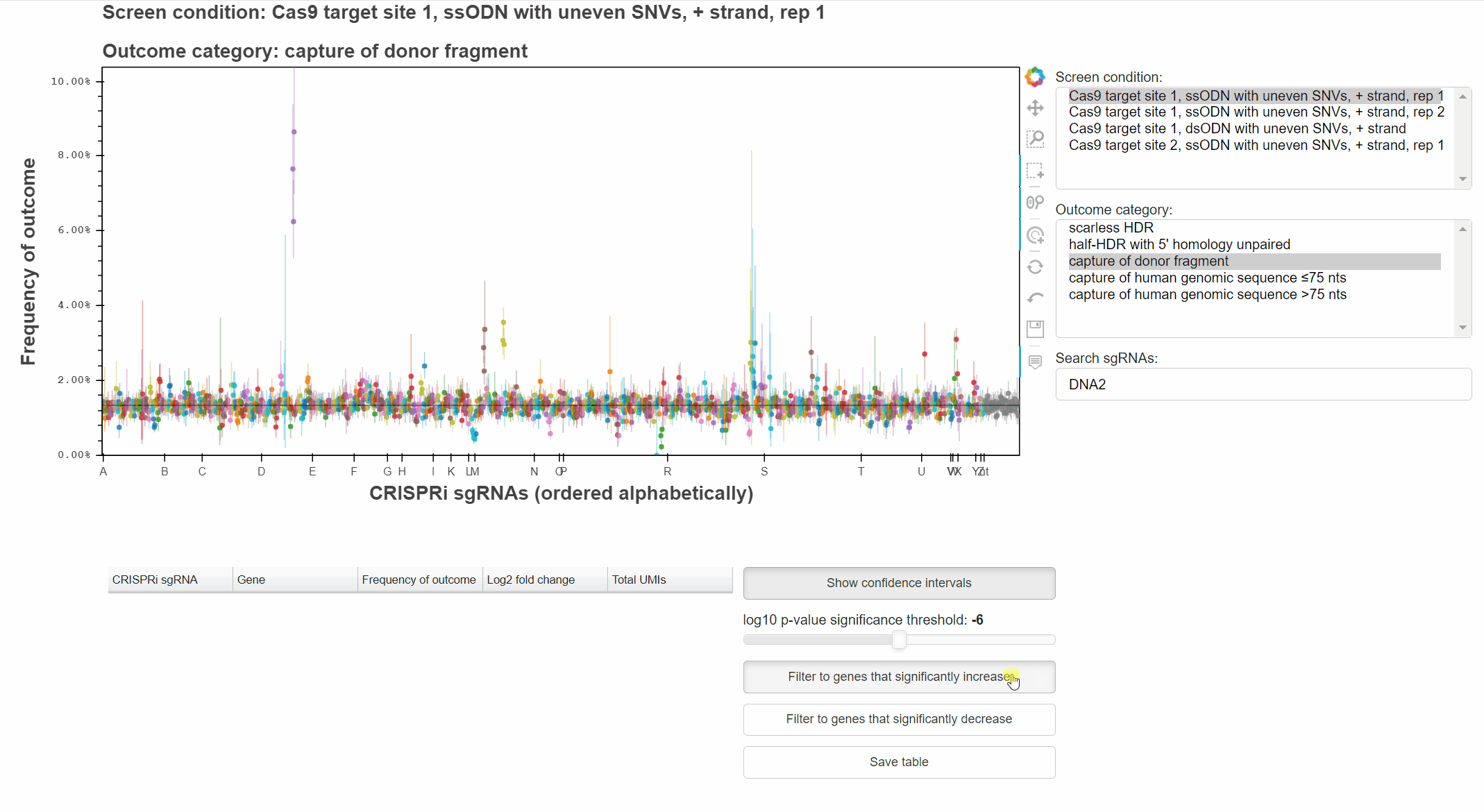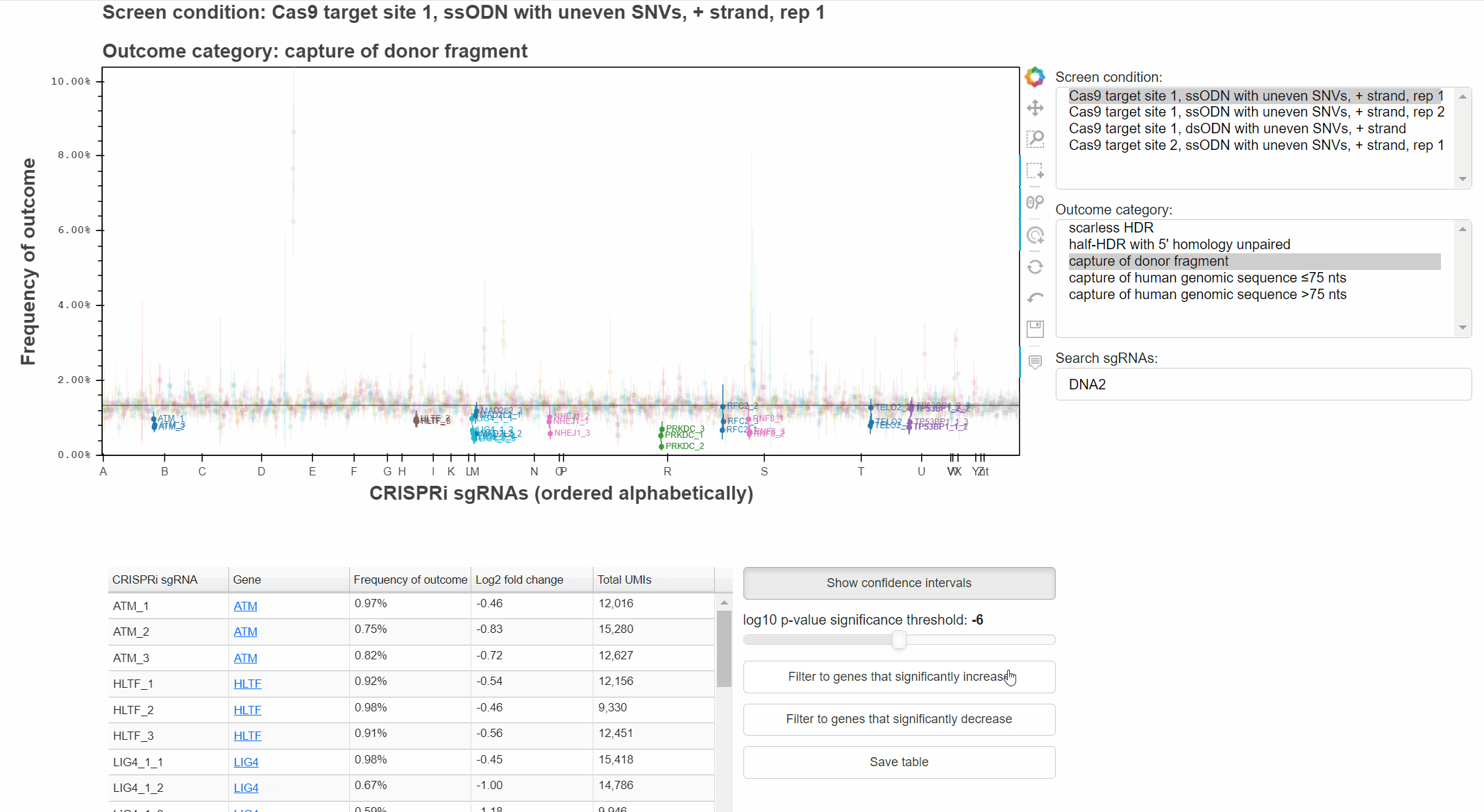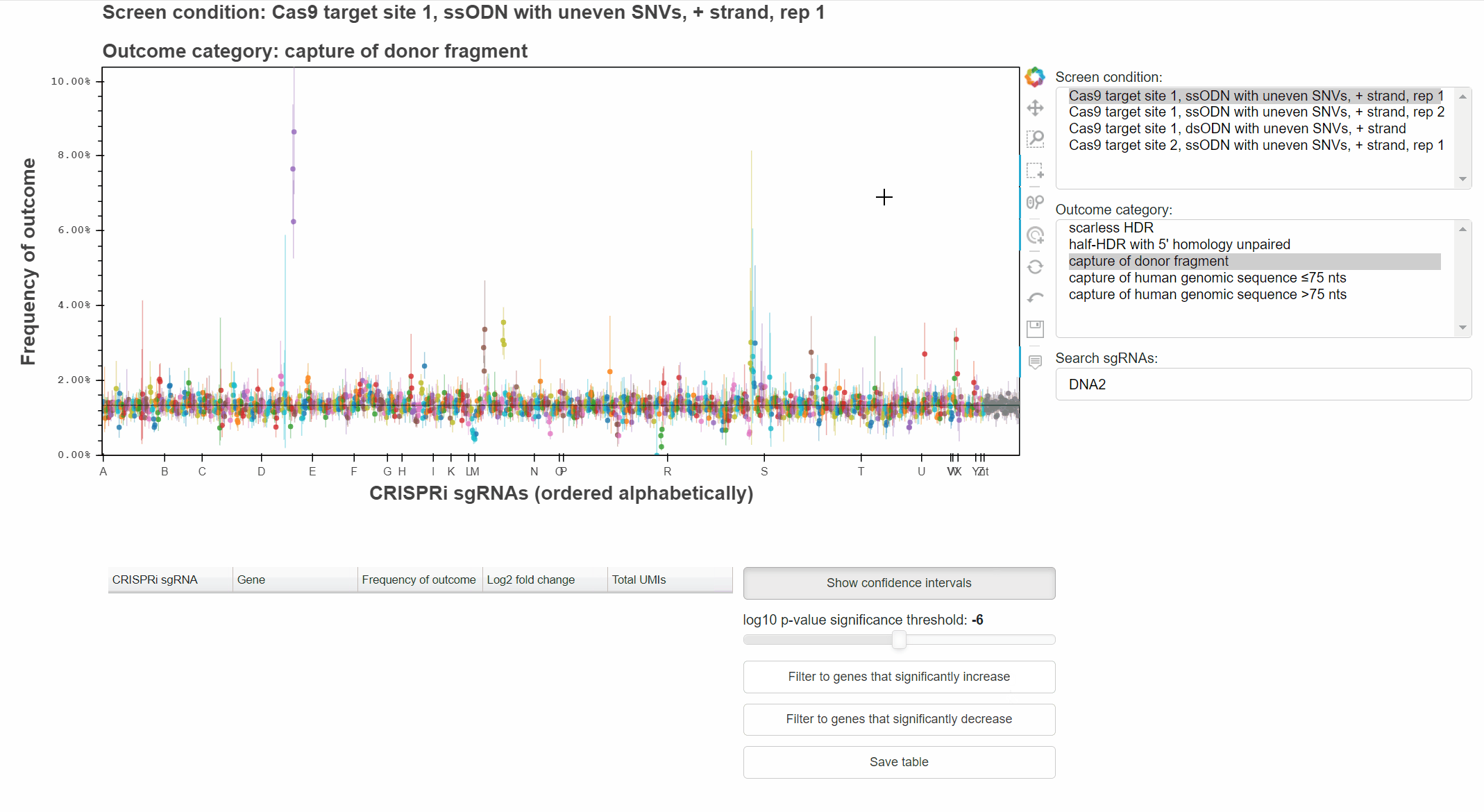
Genes that significantly increase or decrease the frequency of an outcome can be identified by approriately combining relevant p-values for each sgRNA targeting the gene.
The "Filter to genes ..." buttons selects all sgRNAs whose gene-level p-value is below the selected significance threshold for the relevant effect direction.
With the "Wheel zoom" tool active and the cursor over the plot or either axis, scroll with the mouse wheel to zoom.
Plots made using Bokeh.